Journal of Clinical Microbiology and Biochemical Technology
Association of interferon-gamma release assay and SARS-CoV-2 reverse transcriptase polymerase chain reaction test results in adults tested in a tertiary medical center
Evelina N Lagamayo1, Grace C King2* and Ma Concepcion Osano3
2Pathology, Clinical Microbiology, 2nd year Fellow, St. Luke’s Medical Center-Global City, Philippines
3Medical Technologist, Clinical Microbiology, St. Luke’s Medical Center, Philippines
Cite this as
Lagamayo EN, King GC, Osano MC (2021) Association of interferon-gamma release assay and SARS-CoV-2 reverse transcriptase polymerase chain reaction test results in adults tested in a tertiary medical center. J Clin Microbiol Biochem Technol 7(1): 001-005. DOI: 10.17352/jcmbt.000044Introduction: For a country with high Tuberculosis (TB) prevalence, infection of Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) virus with a covert co-infection of Mycobacterium Tuberculosis (MTB) is a real concern. On day 28 symptom onset, he noticed abdominal distension without abdominal pain. CT abdomen revealed pneumoperitoneum. An area of perforation at colon with mucosal haemorrhage was seen in laparotomy. Right hemicolectomy was done. Histology showed mucosal oedema, infiltration with inflammatory cells mainly polymorphs and lymphocytes, congested vessels with areas of haemorrhage in the mucosa.
objective: The study intended to determine if there was an association between the IGRA results and COVID-19 confirmed patients based on their overlapping immunopathophysiology.
Methodology: This was an analytic, cross-sectional study encompassing the two-month enhanced community lockdown in the National Capital Region (NCR) from March 10, 2020 to May 12, 2020. The investigator accessed both the records of IGRA test results with corresponding records of SARS-CoV-2 reverse transcriptase polymerase chain reaction (RT-PCR) test results during the selected study period. Designated code served as patient identifier to protect patient privacy and confidentiality.
Results: The 37 patients included in the study were all adult, mostly in 60-69 age range (mean age 57.5 years) with a slight male predominance (57%). The total detected SARS-CoV-2 cases were 62.2%, whereas the positive IGRA results were 16%. IGRA and SARS-CoV RT-PCR test results have statistically significant association. Majority of COVID-19 confirmed patients exhibited an indeterminate IGRA test result.
Conclusion: IGRA result, being an immune response-dependent test, was susceptible to an on-going SARS-CoV-2 infection due to the similarity of natural host immune response to both MTB and SARS-CoV-2.
Introduction
To analyse the pathogenesis of SARS-CoV-2, in depth studies including genome sequencing and whole array of assays were undertaken. Here, by sheer coincidence of events and timing, the study would attempt to establish an indirect representation of how the virus affects cellular immunity through a routinely run test for tuberculosis. This study hopes to gain an insight into COVID-19 patients’ immune status that may contribute to clinical management.
Around January 22-23, 2020, a novel coronavirus (2019-nCoV) was announced as a Public Health Emergency of International Concern (PHEIC) that eventually developed into a full blown SARS-CoV-2 pandemic around March 11, 2020 [1]. Being an unprecedented occurrence, the Philippine government imposed Enhanced Community Quarantine (ECQ) lockdown particularly in the NCR from March 16 to May 15, 2020 extending to modified ECQ up to May 31, 2020 [2]. This led to hospitals prioritizing admissions to purely emergency cases, a proportion of which are suspected COVID-19 cases. Subsequently, most diagnostic tests run are in-patients; Interferon-gamma Release Assay (IGRA) test in particular was almost exclusively requested for patients exhibiting respiratory symptoms or suspected of tuberculosis infection in our institution. A conspicuous number of indeterminate results in IGRA test prompted an investigation that revealed no technical error and same results on repeat run, the only remarkable similarities being most of the patients were either suspected or confirmed COVID-19 cases.
Interferon Gamma Release Assay (IGRA) is an indirect blood test for tuberculosis infection that measures the in vitro Cell-Mediated Immune (CMI) response against antigens derived from Mycobacterium Tuberculosis Complex (MTC) [3]. Interferon gamma (IFN-ɣ) is produced by CD4+ and CD8+ T lymphocytes and Natural Killer (NK) cells; it plays in important role in inducing and modulating innate and adaptive immune response [4,5]. T-cell release of IFN-ɣ in IGRA is detected by Enzyme-Linked Immunosorbent Assay (ELISA) from blood stimulated with early secreted antigenic target 6 (ESAT-6) and culture filtrate protein 10 (CFP-10), the Mycobacterium Tuberculosis (MTB)-specific antigens [6]. This test is an alternative for Tuberculin Skin Test (TST) that aids in the detecting Latent Tuberculosis Infection (LTBI) and Tuberculosis (TB) disease in conjunction with risk assessment and other medical and diagnostic evaluations. The peptide antigens used are not encoded in the genomes of BCG vaccine strains or most Nontuberculous Mycobacteria (NTM) except M. marinum, M. kansasii, and M. szulgai, making the test more specific than TST [7]. T-SPOT.TB test (Oxford Diagnostic Laboratories, Memphis, TN), the QuantiFERON-TB Gold In-Tube (QFT-GIT; Qiagen, Germantown, MD) test, and most recently, the QuantiFERON-TB Gold Plus (QFT-Plus; Qiagen) assay are the three U.S. Food and Drug Administration (FDA) approved IGRAs as of June 2017 [8], the latter of which is currently used in our institution.
QuantiFERON-TB Gold Plus (QFT-Plus; Qiagen) uses whole blood either preserved in lithium heparin or transferred directly into the four QFT-Plus tubes (Nil, TB1, TB2 and Mitogen tubes). Nil tube contains heparin alone to adjust for background in case of excessive circulating IFN-ɣ or presence of heterophile antibodies; TB1 and TB2 tubes contains peptide antigens ESAT-6 and CFP-10, with TB1 designed to elicit response from CD4+ T helper lymphocytes and TB2 targets response from CD8+ T lymphocytes; mitogen tube serves as the positive control per sample to indicate immune status and check blood handling and incubation. Final results are calculated and interpreted as positive, negative or indeterminate based on the manufacturer’s criteria [9].
This test is particularly helpful in TB endemic countries like the Philippines who hold the third highest prevalence of active tuberculosis in the world after South Africa and Lesotho [10]. With the recent outbreak of SARS-CoV-2, concern for COVID-19 infection among healthcare workers in TB wards and TB patients had been raised [11]. This concern is worsened by the fact that both diseases have a propensity to target the lungs and manifest as pulmonary symptoms. Being both obligate intracellular infectious agents, the natural course of infection would initiate activation of innate immune response to remove and/or neutralize the microbes. Following ingestion by alveolar macrophage for MTB and viral entry into alveolar epithelium for SARS-CoV-2, the host cell would undergo pyroptosis or lytic cell death to expel intracellular pathogens [12,13]. Neighboring epithelial cells, endothelial cells and alveolar macrophages would recognize both the released Damage-Associated Molecular Patterns (DAMPs) and the conserved molecular motif known as Pathogen-Associated Molecular Pattern (PAMP) unique to each organism through the Pathogen Recognition Receptors (PRRs) like Toll-like receptors [14-16]. This will lead to a signaling cascade that activates the transcription factor NFκB, driving the production of pro-inflammatory cytokines (including tumor necrosis factor (TNF), Interleukin-1β (IL-1β), IL-12, IL-6, granulocyte-colony stimulating factor (G-CSF), Monocyte Chemoattractant Protein-1 (MCP-1), Macrophage Colony Stimulating Factor (M-CSF) and interferon gamma-induced protein-10) that will in turn stimulate neighboring T and NK cells to produce additional cytokines like IFN-ɣ [17].
Whereas pulmonary TB would often take an indolent course, COVID-19 patients can either be asymptomatic, moderate or severe acute respiratory syndrome depending on the immune response of host. In healthy individuals, the initial inflammation would attract T cells and macrophages to prevent virus spread, neutralize antibodies, and eliminate virus by phagocytosis with minimal lung damage and eventual recovery. In a dysfunctional/deregulated immune response, the pro-inflammatory feedback loop initiated may cause accumulation of immune cells to target site and unrestrained inflammation known as cytokine storm that mediates widespread lung inflammation and damage on top of direct damage from replicating virus [13].
The point at which innate host response against TB and SARS-CoV-2 intersects is of interest. This study would like to determine if immune response of COVID-19 patients may affect IFN-ɣ release in the peripheral blood samples. If so, this would imply a physiologic response that may render valuable insight to the clinical course of COVID-19 patients potentially co-infected with TB. As of writing, no study of similar nature has been published.
Objectives
General objective: To determine the association of SARS-CoV-2 RT-PCR and IGRA test results in adult patients tested for both diagnostic examinations.
Specific objectives
1. To determine the proportion of patients tested for COVID-19 who have positive results for SARS-CoV-2 RT-PCR.
2. To determine the proportion of patients tested for COVID-19 who have positive results for IGRA.
3. To determine the association of SARS-CoV-2 RT-PCR and IGRA test results.
Methods
This was an analytical, cross-sectional study that included all IGRA results with corresponding SARS-CoV-2 test results from March 10, 2020 to May 12, 2020.
IGRA tests were routinely run on Tuesdays except on STAT cases. The study period selected started from the second week of March until the second week of May2020, encompassing the start to end of enhanced community quarantine in NCR. The investigator accessed both the institution’s Clinical Microbiology hard copy records of IGRA test results and Clinical Immunology’s laboratory information system records of SARS-CoV-2 test results during the selected study period. Designated code served as patient identifier to protect patient privacy and confidentiality.
Inclusion criteria
All IGRA test results and their corresponding RT-PCR SARS-CoV-2 test results run during the study period were included. In case of repeat testing for RT-PCR SARS-CoV-2, the closest date between the time of specimen collection for both IGRA and RT-PCR SARS-CoV-2 tests were chosen. The maximum date for RT-PCR SARS-CoV-2 test included was 15 days from the extraction date for the IGRA test based on the symptom onset range derived by Lauer, et al. (2020) [18].
Exclusion criteria
• Duplicate results (if any).
• IGRA test results without RT-PCR SARS-CoV test results
Operational definitions
• IGRA: 1 ml blood was placed into each of the four QFT-Plus blood collection tubes, incubated, and centrifuged to separate and harvest the plasma. This was then processed according to the manufacturer’s instructions. The optical density was measured and calculated using the QFT-Plus Analysis Software for result interpretation.
• Positive IGRA: IFN-ɣ response to either TB Antigen tube was significantly above the Nil IFN-ɣ IU/ml value, so M. tuberculosis infection was likely
• Negative IGRA: IFN-ɣ response to either TB Antigen tube was not significantly above the Nil IFN-ɣ IU/ml value, so M. tuberculosis infection was NOT likely
• Indeterminate IGRA: there was either a low response to Mitogen or the Nil values maybe outside the assay measurement range, so the likelihood of M. tuberculosis infection cannot be determined
• RT-PCR for SARS-CoV-2: Nasopharyngeal and oropharyngeal swabs placed in a viral transport media underwent SARS-CoV-2 RNA extraction by reverse transcription and processed according to the manufacturer’s recommendation. Amplification reactions were monitored using fluorescence and computed by equipment-specific analysis software. Presence or absence of target RNA was reported as DETECTED or NOT DETECTED after a final validation by a consultant Pathologist.
Sample size estimation
Assuming a level of significance of 0.05, a power of 80%, a degree of freedom of 2, and a medium effect size (0.3), the required sample size to run a Chi-Square Test of Association is 108.
Data analysis
Data were analysed using proportion expressed in percentages using the following formulae:
% SARS-CoV-2 RT PCR detected = (No. of DETECTED RT-PCR/Total number of samples) x 100%
% Positive IGRA = (No. of POSITIVE IGRA/Total number of samples) x 100%
The association of positive, indeterminate and negative IGRA results with confirmed SARS-CoV-2 detection were measured by Fisher’s exact test.
Ethical considerations
• The clinical protocol and all relevant documents were reviewed and approved by the SLMC Institutional Ethics Review Committee.
• Patient confidentiality was respected by ensuring anonymity of patient records. Only the designated code and results were recorded and stored by the project leader.
• The manner of disseminating and communicating the study results guaranteed the protection of the confidentiality of patients’ data.
• Electronic data were stored in a password-protected laptop and were only accessible to the project and co-project leaders. The password will be the sole responsibility of the project leader.
• Study-related documents will be stored in a cabinet with lock and key. The key is kept by the project leader and the cabinet, which is located in a secure room, will only be accessible to members of the research team.
• All documents and electronic data will be kept by the project leader for a standard storage period of at least five years after completion of study and shall be securely shredded and/or erased.
Results
Demographic data
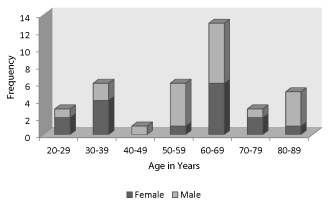
A total of 37 patients were included in the study, 21 (57%) of which were male and of these, 14 (14/23, 60.87%) were COVID-19 confirmed cases. The patients were all adults, the youngest at 21 years old while the oldest at 89 years old. The mean age was 57.5 years old (61.13 years old for confirmed COVID-19 cases; 51.57 years old for non-COVID-19 cases). Most patients were clustered at age range of 60-69 years old, while the least was at 40-49 years old (Figure 1).
Association of IGRA and SARS-CoV-2 RT-PCR test results
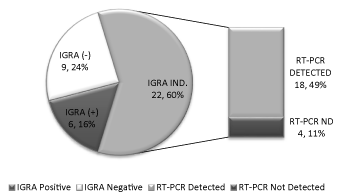
There was a statistically significant association between IGRA and SARS-CoV-2 RT-PCR test results (p=0.006) (Table 1). The total number of detected SARS-CoV-2 cases was 62.2% (23 patients), while the positive IGRA results was 16% (6 patients). Majority of the IGRA results were indeterminate (60%) and where most of the detected SARS-CoV-2 cases were found (49%) Figure 2.
Discussion
Demographic data of the study was similar to reported cases during the initial outbreak where SARS-CoV-2 detected cases were elderly with slight male predominance [19-20]. Notably, most of these detected cases had indeterminate results in IGRA. Indeterminate IGRA results can be attributed to pre-analytical, analytical and immunological sources of variability [6]. Pre-analytical and analytical variability like blood volume, tube shaking, incubation duration and manufacturing defects, although not completely eliminated, were presumed minimized due to the stringent quality control practices in place from specimen receiving to processing. As previously mentioned, the increased indeterminate results prompted a repeat run with additional inclusion of prior known positive specimen to ascertain validity of the previous run. This then leaves immunological variability as the most likely cause of indeterminate results.
Immunological variability may either be due to immune boosting or immunomodulation [6]. Immune boosting can be from excessive levels of circulating IFN-ɣ or the presence of heterophile antibodies; this will be translated computationally as a Nil value higher than the recommended cut-off [9]. Immunomodulation, on the other hand, may occur with insufficient lymphocytes or the functional deficiency of lymphocyte to produce IFN-ɣ; this then will be represented computationally as Mitogen value below the recommended cut-off [9]. In this study, all indeterminate results were of the latter case (data not shown). Majority of positive IGRA results do not progress to active TB disease [6]. In case of SARS-CoV-2 and LTBI co-infection though, the sustained depressed T cell mediated response may increase risk of reactivation of active TB disease.
Studies of the immune response to SARS-CoV-2 virus emphasized peripheral lymphopenia, mainly CD4 and CD8 T cells, associated with symptomatic and/or severe form of the disease [21-25]. Sequestration of cytokine and chemokine-recruited immune cells from the blood into the infected site, particularly the lung; lymphocytic infiltration of the airway; T cell exhaustion; and viral-initiated/induced lymphocytic apoptosis were posited as possible reasons for the lymphopenia [13,21,24,26]. These evidences support the significant association of IGRA indeterminate results and SARS-CoV-2 detected patients, implying inadequate peripheral lymphocytes due to the host immune response’s attempt to achieve an antiviral state through signalling cascades in response to the infection. Simply put, the patients with indeterminate IGRA results and RT-PCR SARS-CoV-2 detected results were probably in the acute phase of symptomatic and/or severe form of COVID-19. In highly prevalent areas where TB infection is a constant threat on top of the existing SARS-CoV-2 pandemic, this test can alert clinicians to expect symptomatic COVID-19 with possible severe course. The study is limited by the sample size and the possibility of viral evolution with corresponding change in host immune response. Further study of the clinical correlation could provide a clearer picture as to whether the deficient peripheral T cell response to IGRA, in turn, proved to be a protective or detrimental response to SARS-CoV-2 virus.
Conclusion
IGRA can serve not only as an indirect tool for LTBI detection, but can also implicitly reflect the state of a patient’s adaptive immune response, particularly the T lymphocytes. Therefore, IGRA can indirectly assess the adequacy of T cell response to two highly infectious pulmonary pathogens, that is, MTB and SARS-CoV-2.
(Appendix)
- Timeline of WHO’s response to COVID-19 (2020). Link: https://bit.ly/3iTgyYB .
- Timeline of the COVID-19 pandemic in the Philippines (2020). Link: https://bit.ly/3a5yQSs
- Mazurek GH, Jereb J, Vernon A, LoBue P, Goldberg S, et al. (2010) Updated Guidelines for Using Interferon Gamma Release Assays to Detect Mycobacterium Tuberculosis Infection --- United States, 2010. Link:https://bit.ly/3pxifh2
- Tau G, Rothman P (1999) Biologic functions of the IFN-gamma receptors. Allergy 54: 1233-1251. Link: https://bit.ly/2YnBNZ2
- Miller CH, Maher SG, Young HA (2009) Clinical Use of Interferon-gamma. Ann N Y Acad Sci 1182: 69-79. Link: https://bit.ly/2Ynf1Ra
- Pai M, Denkinger CM, Kik SV, Rangaka MX, Zwerling A, et al. (2014) Gamma interferon release assays for detection of Mycobacterium Tuberculosis infection. Clin Microbiol Rev 27: 3-20. Link: https://bit.ly/39rDpXW
- Andersen P, Munk ME, Pollock JM, Doherty TM (2000) Specific immune-based diagnosis of tuberculosis. Lancet 356: 1099-1104. Link: https://bit.ly/3aa44rt
- Theel ES, Hilgart H, Breen-Lyles M, McCoy K, Flury R, et al. (2018) Comparison of the QuantiFERON-TB Gold Plus and QuantiFERON-TB Gold In-Tube Interferon Gamma Release Assays in Patients at Risk for Tuberculosis and in Health Care Workers. J Clin Microbiol 56: e00614- e00618. Link: https://bit.ly/3a7DSxW
- Qiagen (2016) QuantiFERON® - TB Gold Plus (QFT®-Plus) ELISA.
- Weiler, Gundo Aurel. (2019, March 24). It’s time to end TB in the Philippines. Link: https://bit.ly/3af565y
- Sayo AR, Balinas EGM, Verona JA, Villanueva AMG, Han SM, et al. (2020) COVID-19 screening on a tuberculosis ward in Manila, the Philippines. J Clin Tuberc Other Microbact Dis 20. Link: https://bit.ly/3j0WW4P
- Jorgensen I, Miao EA (2015) Pyroptotic cell death defends against intracellular pathogens. Immunol Rev 265: 130-142. Link: https://bit.ly/3ch7vPM
- Tay MZ, Poh CM, Rénia L, MacAry PA, Ng LFP, et al. (2020) The trinity of COVID-19: immunity, inflammation and intervention. Nat Rev Immunol 20: 363-374. Link: https://bit.ly/39pHcFx
- Stamm CE, Collins AC, Shiloh MU (2015) Sensing of Mycobacterium tuberculosis and consequences to both host and bacillus. Immunol Rev 264: 204-219. Link: https://bit.ly/2Yp8aGV
- Li X, Geng M, Peng Y, Meng L, Lu S (2020) Molecular immune pathogenesis and diagnosis of COVID-19. J Pharm Analysis 10: 102-108. Link: https://bit.ly/3j0XpE7
- Totura AL, Baric RS (2012) SARS coronavirus pathogenesis: host innate immune responses and viral antagonism of interferon. Curr Opin Virol 2: 264-275. Link: https://bit.ly/3sY5iPn
- Mutar Mahdi B (2020) COVID-19 type III hypersensitivity reaction. Med Hypotheses 140: 109763. Link: https://bit.ly/3rbFgXh
- Lauer SA, Grantz KH, Bi Q, Jones FK, Zheng Q, et al. (2020) The Incubation Period of Coronavirus Disease 2019 (COVID-19) From Publicly Reported Confirmed Cases: Estimation and Application. Ann Intern Med 172: 577-582. Link: https://bit.ly/3sXblDE
- Chen N, Zhou M, Dong X, Qu J, Gong F, et al. (2020) Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet 395: 507-513. Link: https://bit.ly/3pmIJBA
- Yuki K, Fujiogi M, Koutsogiannaki S (2020) COVID-19 pathophysiology: A review. Clin Immunol 215: 108427. Link: https://bit.ly/3pqcDFf
- Huang C, Wang, Yeming W, Li X, Ren L, et al. (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395. Link: https://bit.ly/3j1yrVi
- Li H, Liu L, Zhang D, Xu J, Dai H, et al. (2020) SARS-CoV-2 and viral sepsis: observations and hypotheses. Lancet 395: 1517-1520. Link: https://bit.ly/3ordl3q
- Huang I, Pranata R (2020) Lymphopenia in severe coronavirus disease-2019 (COVID-19): systematic review and meta-analysis. J Intensive Care 8: 36. Link: https://bit.ly/3t30AQm
- Jesenak M, Brndiarova M, Urbancikova I, Rennerova Z, Vojtkova J, et al. (2020) Immune Parameters and COVID-19 Infection – Associations With Clinical Severity and Disease Prognosis. Front Cell Infect Microbiol 10: 364. Link: https://bit.ly/3pDpEuU
- Zhou Y, Fu B, Zheng X, Wang D, Zhao C, et al. (2020) Pathogenic T cells and inflammatory monocytes incite inflammatory storm in severe COVID-19 patients. Natl Sci Rev nwaa041. Link: https://bit.ly/3om24Bv
- Huang KJ, Su IJ, Theron M, Wu YC, Lai SK, et al. (2005) An interferon-gamma-related cytokine storm in SARS patients. J Med Virol 75: 185-194. Link: https://bit.ly/39lZO9g

Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.
 Save to Mendeley
Save to Mendeley
