Archives of Otolaryngology and Rhinology
Profiling ACE2 and TMPRSS2 expression in sinonasal mucosa
Michael E Price1, Rohit Nallani1, Jared S Bodine1, Gunjan Gaur1, Levi Arnold1, Bryan Humphrey1, Brendan Ottemann1, Maura F O’Neil2, Kevin J Sykes1, D David Beahm1, Jianming Qiu3, Alexander G Chiu11**, Sufi Mary Thomas1*
2Department of Pathology and Laboratory Medicine, University of Kansas Medical Center, Kansas City, KS, USA
3Department of Microbiology, Molecular Genetics and Immunology, University of Kansas, Kansas City, KS, USA
Cite this as
Price ME, Nallani R, Bodine JS, Gaur G, Arnold L, et al. (2022) Profiling ACE2 and TMPRSS2 expression in sinonasal mucosa. Arch Otolaryngol Rhinol 8(4): 020-026. DOI: 10.17352/2455-1759.000148Copyright
© 2022 Price ME, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.Rhinologists may be one of the highest-risk subspecialties in otolaryngology for exposure to SARS-CoV-2 as the sinonasal passage seems to be a reservoir for the virus. Previous data indicate nasal epithelial cells express the primary receptor for SARS-CoV-2, Angiotensin-Converting Enzyme-2 (ACE2). However, no data exist profiling the regional expression of ACE2 or the expression of transmembrane serine protease 2 (TMPRSS2), an additional protease necessary for SARS-CoV-2 viral entry, within the sinonasal cavity. We sought to assess for anatomic expression of ACE2 and TMPRSS2 throughout the nasal cavity and paranasal sinuses. We hypothesize that ACE2 and TMPRSS2 are expressed throughout the nasal cavity and paranasal sinuses.
To test this hypothesis, we sampled various regions of the sinonasal cavity from patients undergoing rhinology procedures and used immunohistochemical staining to profile ACE2, compare ACE2 expression between regions, and compare ACE2 expression between patients and patient characteristics.
We found ACE2 and TMPRSS2 are present throughout the sinonasal cavity without a regional pattern among anatomic regions in our patients. We found no statistically significant correlation in ACE2 expression with patient characteristics such as age, sex, or BMI. We also did not find a statistically significant correlation between ACE2 and TMPRSS2 quantitative expression. ACE2 expression trended higher in males compared to females for six out of seven regions excluding the nasal floor.
In conclusion, ACE2 and TMPRSS2 are expressed ubiquitously throughout the sinonasal cavity. ACE2 expression may be higher in the sinonasal cavity in males compared to females. These data implicate that SARS-CoV-2 is unlikely to discriminate between anatomic regions as a point of entry and that anatomic regions likely are similar in viral load. Thus, all rhinology and skull base surgeries, independent of encounter of the anatomic region in the sinonasal cavity, predicate screening for SARS-CoV-2, and necessary personal protective equipment.
Introduction
The SARS-CoV-2, Coronavirus Disease (COVID-19) pandemic has triggered a global emergency. To date, over 3 million cases of COVID-19 have been diagnosed worldwide. The healthcare industry is focused on creating operating principles that protect both patients and providers who may be exposed during their care of COVID-19 symptomatic and asymptomatic patients. Early reports indicate some specialties are more at risk for contracting COVID-19. Working in proximity to anatomic regions high in viral load and performing aerosol-generating procedures may expose anesthesiologists, pulmonologists, and otolaryngologists at a higher rate than other physician specialties. Rhinologists, or otolaryngologists who specialize in endoscopic sinus and skull base surgery, may be one of the highest-risk subspecialties in otolaryngology. An endoscopic, pituitary surgery in Wuhan, China has been reported to have infected fourteen individuals involved in case [1]. This has since been challenged by others who believe the infections happened in the post-operative care of the patient [2]. Regardless, the actual risk of performing these types of procedures is currently unknown.
The nasal cavity is a common site of viral entry, and viral loads in symptomatic patients are higher in the nose than in the throat [3,4]. However, the viral load within the paranasal sinuses, in comparison to the nasal cavity, nasopharynx, or oropharynx, is also currently unknown. Membrane-associated Angiotensin-Converting Enzyme-2 (ACE2) serves as the primary receptor for SARS-CoV-2 [5]. Before cell entry, SARS-CoV-2 must be cleaved by transmembrane serine protease 2 (TMPRSS2). Research on SARS-CoV, a genetic relative of SARS-CoV-2, following the SARS epidemic showed infection of nasal and tracheobronchial cells is ACE2 receptor-dependent and can result in the shedding of infected ciliated cells into the airway lumen [6]. Recent data corroborate a similar entry mechanism for SARS-CoV-2 [7]. Identifying anatomic regions rich in ACE2 and the necessary TMPRSS2 may serve to aid in determining the risk of viral load exposure during endoscopic or skull base surgery. On top of determining surgeons’ risk of contracting the viral illness, an analysis of patient characteristics associated with higher levels of ACE2 and TMPRSS2 may open doors to understanding which patients have a higher risk for earlier symptom onset, more severe disease, or increased transmissibility. The role of the relationship of ACE2 and TMPRSS2 expression with asthma and allergic sinusitis/rhinitis is still developing and may play an important role in both inpatient risk of COVID-19 and surgeon risk of exposure to SARS-CoV-2 in rhinology surgery. Additionally, a comparison of ACE2 expression across epithelial tissue of patients with inflammatory pathologies such as cancer could shed light on the role of ACE2 in diseases besides COVID-19.
The distribution and regional expression of ACE2 and TMPRSS2 throughout the nasal cavity and paranasal sinuses are unknown. We hypothesize that ACE2 and TMPRSS2 are co-expressed throughout the nasal cavity and paranasal sinuses. To test this hypothesis, we sampled nasal mucosa from patients undergoing rhinologic surgery and assessed for ACE2 and TMPRSS2 expression via immunohistochemistry. Recently, the olfactory epithelium was shown to express ACE2 at a markedly higher rate than non-olfactory epithelium [8]. We sought to map the relative ACE2 and TMPRSS2 expression throughout the latter tissue within anatomic regions of the nasal cavity and paranasal sinuses. We also compared ACE2 and TMPRSS2 levels across patient characteristics. Understanding the relative burden of disease within the sinuses may help guide physicians in determining their personal risk of contracting COVID-19 as well as long-term considerations in wound healing and outcomes following sinus procedures in COVID-19-positive patients. Finally, a correlation of ACE2 expression with cancer survival times aims to expand the discussion of the enzyme to include other disease processes.
Methods
Patients and clinical specimens
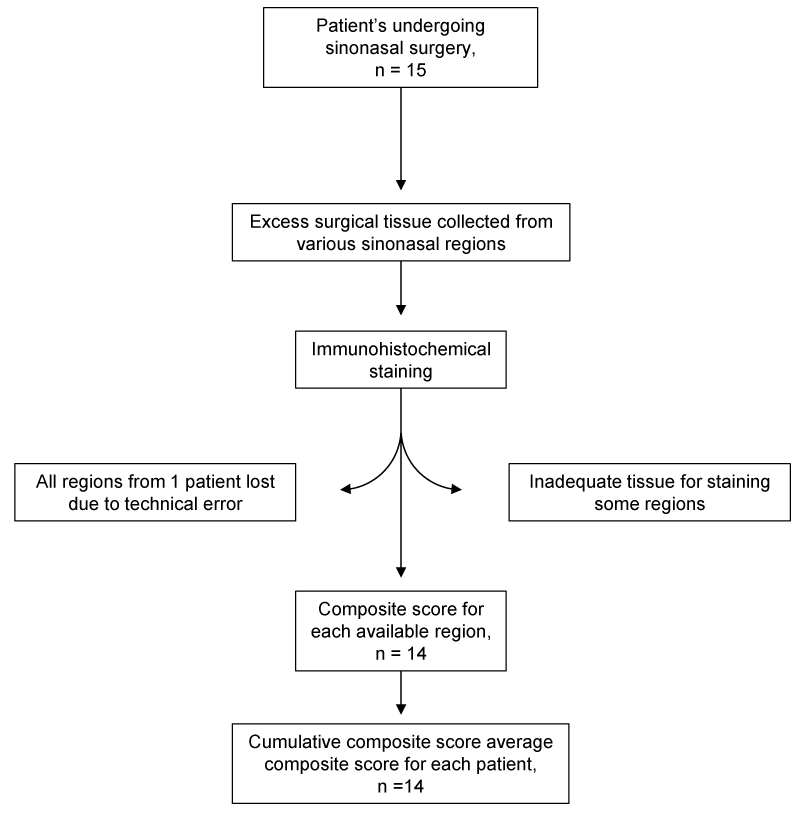
Formalin-Fixed Paraffin-Embedded (FFPE) tissue specimens from fifteen patients were collected by the Department of Otolaryngology, University of Kansas Medical Center. All patients were either screened with a University of Kansas Hospital surgical screening protocol or by COVID-19 viral testing and as well as PCR. All patients tested negative with no known history of COVID-19-positive testing. The Institutional Review Board approved the study after a review of compliance with ethical standards and samples were collected and deidentified after obtaining written informed consent. Nasal mucosal tissue from seven different regions within the nasal cavity (nasal floor, sphenoid sinus, inferior turbinate, ethmoid bulla, middle turbinate, uncinate process, and nasopharynx) was collected from patients undergoing procedures that resulted in excess surgical tissue of these regions.
Immunohistochemical analysis
Sections (4µm) from formalin-fixed, paraffin-embedded samples mounted on adhesive slides were subjected to immunohistochemical staining of ACE2. Briefly, after deparaffinization and rehydration, tissue sections were pre-treated using citrate buffer for 5 min in a pressure cooker. After 10 min cooldown, hydrogen peroxide (3%) was applied to the sections to quench endogenous peroxidase activity. Sections were then incubated with primary antibodies against ACE2 (rabbit polyclonal ab15348; 1:2000 dilution; Abcam, Cambridge, MA) or TMPRSS2 (rabbit polyclonal PA5-83286, ThermoFisher, Waltham, MA) for 30 min., buffer rinsed and followed by a 30 min incubation with MACH 2TM Rabbit HRP-Polymer (Biocare Medical, Pacheco, CA, USA). Finally, the staining was visualized by DAB+ (Dako, Carpinteria, CA, USA). Immunohistochemical staining was performed using the Biocare Medical IntelliPATH FLX Automated Stainer at room temperature. A light hematoxylin counterstain was performed, following which the slides were dehydrated, cleared, and mounted using permanent mounting media.
Aperio ImageScope (Version 12.3.0) was used for the computer-based analysis of digitally scanned slides. Epithelial cell compartments were cordoned off digitally from stromal compartments using a circle tool. ACE2 expression was quantified using the cytoplasmic analysis function and TMPRSS2 expression was quantified using the sum of the cytoplasmic and nuclear analysis function. Percent cytoplasmic staining for each intensity of the stain on the +1, +2, or +3 scale was determined. Composite scores were calculated by multiplying the percent positivity with the intensity score. The composite scores at each intensity were then added for each slide to get the total composite score. The cumulative composite score was the average composite score of all regions sampled per patient. Statistics were performed using GraphPad Prism by non-parametric testing with either Mann Whitney or one-way ANOVA with Kruskal-Wallis post-test as indicated in the text and figure legends.
PCAWG analysis
Pan-Cancer Analysis of Whole Genomes (PCAWG) was used to determine predicted cancer patient outcomes in high/low ACE2 expression. The log-rank test validated the significance between the high and low groups. Kaplan-Meier plots were generated and downloaded from the PCAWG Xena browser.
PCAWG pan-cancer data for ACE2 and TMPRSS2 were downloaded from the Xena browser using the PCAWG donor-centric cohort. Datasets were then uploaded to GraphPad Prism 8 and Pearson correlation plot was generated (r > 0.4 significance cutoff).
Results
ACE2 and TMPRSS2 expression and patient characteristics
We included samples from fifteen patients in our study. The patients included were 14 adults with a median age of 56.5 years and a range of 23-86 years and one pediatric patient. The sex distribution was 22% female (4/15). 11 patients were never smokers, three patients were former smokers, and one patient was a current smoker. None of the patients had Angiotensin-Converting Enzyme (ACE) inhibitors or angiotensin receptor blockers as medications listed in their medical history. To determine if there was any correlation between ACE2 and TMPRSS2 expression with patient characteristics we compared regional composite scores and cumulative positive scores with patient characteristics (Table 1). There was no statistical correlation between ACE2 or TMPRSS2 composite scores and any of the patient characteristics including comparing patients with sinusitis compared to patients without sinusitis (51.38 ± 10.78 for control cumulative composite score; n = 9 compared to 48.52 ± 23.12 composite score for sinusitis; n = 5; p > 0.9999 by Mann-Whitney test) Figure 1. There was, however, a trend for males to have higher regional ACE2 in all seven regions sampled and a trend toward a higher average ACE2 composite (Figure 2A-B). All patients from whom TMPRSS2 was quantified were male.
ACE2 and TMPRSS2 are expressed throughout various regions of the nasal cavity and localize primarily to the epithelium
We hypothesized that nasal cavity mucosal ACE2 expression would vary based on region. We found positive ACE2 expression across all sites sampled from every patient (Figure 3A-N) except for one sphenoid sinus sample (Figure 3A-N). Within the mucosa, ACE2 localized primarily to the cytoplasm of cells, independent of the site sampled (Figure 3A-N). The single sample that did not demonstrate ACE2 staining was devoid of epithelium, most likely due to an error in sampling or processing (data not shown). Similarly, TMPRSS2 was expressed throughout every sample tested in every region (Figure 3O-U).
ACE2 and TMPRSS2 expression varies between patients and anatomic region of the nasal cavity and paranasal sinuses
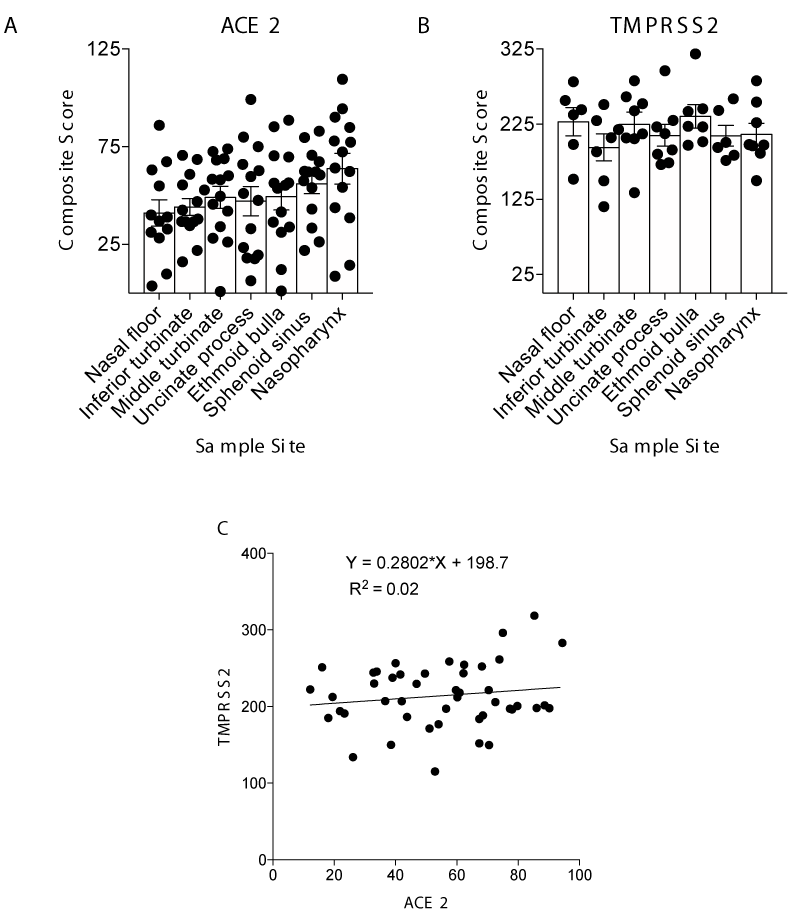
To further understand the relative expression in each region, we quantified both percent positivity and signal intensity to develop a composite score (percent positivity multiplied by intensity) for each region. Regional ACE2 and TMPRSS2 expression varied between regions and among patients (Figures 4A and 4B). There was no statistically significant difference in composite scores between different regions among all the patients for ACE2 or TMPRSS2 (Figure 4A; p = 0.22 by Kruskal-Wallis test; Figure 4B; p = 0.54 by Kruskal-Wallis test). However, the nasopharynx trended toward higher ACE2 expression, and the nasal floor trended toward lower ACE2 expression. Finally, while the presence of both ACE2 and TMPRSS2 was ubiquitous in all samples evaluated, there was no statistically significant correlation between ACE2 and TMPRSS2 composite score (Figure 4C; R2 = 0.02, by Pearson Correlation linear regression). In combination, these data suggest no generalized regional specificity of ACE2 or TMPRSS2 expression within the nasal cavity.
Pan-cancer analysis of whole genomes (PCAWG) expression
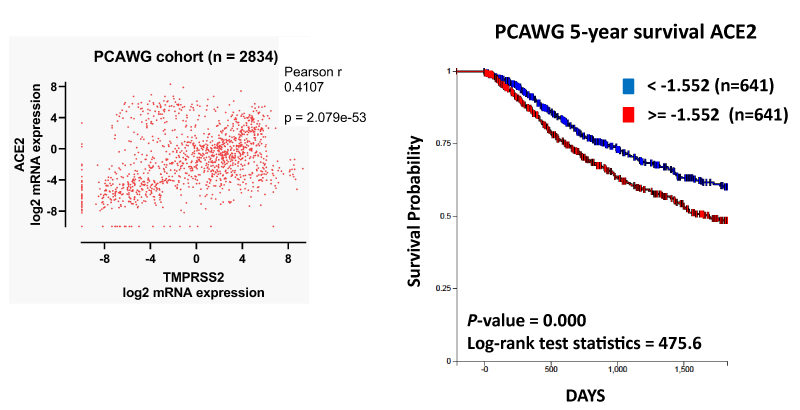
To determine the relevance of ACE2 and TMPRSS2 expression in cancer pathologies, we utilized the PCAWG dataset, a project combining the International Cancer Genome Consortium (ICGC) and The Cancer Genome Atlas (TCGA) datasets across 38 tumor types (n = 2,834). We compared ACE2 and TMPRSS2 mRNA expression and found that, at a Pearson cutoff of > 0.4, the two genes were significantly correlated in the dataset (Figure 5A). We then evaluated the 5-year survival of cancer patients who were either high or low ACE2 expressors. We found that high ACE2 expressors showed a significant decrease in survival probability compared to low ACE2 expressors (Figure 5B). There were 641 patients in both high and low groups.
Discussion
Our data demonstrate that ACE2 and TMPRSS2 are ubiquitously present throughout the epithelium of various regions of the nasal cavity, paranasal sinuses, and nasopharynx. Importantly, relative ACE2 and TMPRSS2 regional expression vary widely between patients and there is no consistent pattern of ACE2 or TMPRSS2 relative expression to a specific region. The presence of ACE2, the receptor for SARS-CoV-2, and the protease, TMPRSS2, throughout these regions highlights the risk of viral exposure when performing procedures that access these regions. Because ACE2 and TMPRSS2 proteins are more relevant than mRNA for viral entry, mRNA levels were not quantified.
Clinical sampling supports the nasal cavity to harbor SARS-CoV-2 viral load. Strong evidence supports ACE2 as the receptor for entry of the SARS-CoV-2 virus [7]. Our data corroborate previous reports of ACE2 expression in the paranasal sinuses [9]. Other data suggest the relative abundance of ACE2 in the upper respiratory tract is less than that of the lower respiratory tract [10]. This relatively lower ACE2 presence in the upper respiratory tract has been suggested to limit the transmissibility of SARS-CoV [11]. In the context of SARS-CoV-2, a report of two patients demonstrated comparable viral load from throat swabs compared to sputum samples [12]. A separate study demonstrated higher viral load from nasal swabs compared to throat [3,4]. In combination, data from these reports suggest the viral load in the nasal cavity is similar, if not greater, than that of the lower airway.
Extrapolation of ACE2 localization to viral load in the nasal cavity and paranasal sinuses, and risk of contracting the virus during procedures that access these regions is limited. Recent data suggest that while necessary for viral entry, ACE2 alone is not sufficient for viral infectivity [7]. Co-expression of the serine protease TMPRSS2 or a cathepsin is apparently necessary for viral infectivity and replication [7]. Thus, presence of ACE2 alone may not be the sufficient to characterize regional viral burden potential within a tissue. Despite this, it is well established that the nasal cavity is an abundant source of viral load, and the epithelial cells of the nasal cavity likely harbor the molecular machinery necessary for viral entry and replication. Although our samples were derived from a small yet diverse group of patients, limiting matching of our samples, our data clearly indicate that ACE2 is expressed throughout the nasal cavity. However, the inter-patient and inter-region variability of ACE2 expression in a small sample size, makes it difficult to generate statistical confidence related to regional ACE2 expression. Identifying potential confounders of ACE2 expression in the nasal cavity and paranasal sinuses or sampling from a larger more homogenous group of patients may allow for better mapping of the anatomical distribution of ACE2. This, however, may be of little benefit as sinonasal disease or SARS-CoV-2 infection may alter ACE2 expression. To further elucidate the relative contribution of specific anatomic regions to viral burden, direct viral measurement from infected patients is needed.
Our data identify sex as a potential source of variability of ACE2 expression (Figure 2A-B) and the trend for male ACE2 to be higher, although not statistically significant, warrants further evaluation. Early epidemiological reports showed that male individuals were more commonly affected, although more recent reports demonstrate equivalent distribution of infected cases [13-15]. Whether or not sex is a risk factor for SARS-CoV-2 transmissibility is unknown, and it may be worthwhile in future studies to investigate whether the trend toward higher ACE2 expression in men has implications for increased transmissibility, higher severity, or earlier onset of disease.
Additional risk factors for SARS-CoV-2 disease severity may also be associated with ACE2 expression or viral load in the nasal cavity. Due to our small sample size and heterogeneous cohort, we did not identify any relationship between other factors such as age, comorbidities, or smoking status. Moreover, our study was designed to serve as a preliminary anatomic atlas of ACE2 and TMPRSS2 expression and was not powered to attempt to identify differences driven by patient characteristics. These secondary analyses serve only as preliminary data to support the need for additional study and should be interpreted with caution. Our data are also limited by ethnicity as all but one of our patients selected white non-Hispanic as the ethnic designation. Our study group consisted only 2 former smokers and one current smoker limiting any conclusion about the role of smoking in ACE2 and TMPRSS2 expression. Analyses of cancer genome databases suggests that ACE2 expression correlates with reduced survival in cancer patients, drawing a potential biomolecular link between cancer and COVID-19 and highlighting the role of ACE2 in pathologies associated with inflammation. This finding prompts further study into a mechanistic understanding of the potential role of ACE2 in malignancy. The mechanistic relationships possibly linking mortality of ACE2, TMPRSS2 and SARS-CoV2 are beyond the scope of this study and more work is needed in this area.
In summary, we have demonstrated that ACE2, the SARS-CoV-2 receptor, and TMPRSS2, a protease necessary for SARS-CoV-2 viral entry, are present throughout the epithelium of the nasal cavity and paranasal sinuses. ACE2 nor TMPRSS2 expression does not have apparent anatomic patterns but ACE2 may be increased in the nasal cavity and paranasal sinuses of males compared to females. These data reinforce the need for caution, screening for SARS-CoV-2, and use of necessary protective equipment when performing rhinologic and skull base surgeries during this time.
Financial support
The University of Kansas Cancer Center under CCSG P30CA168524, NIH grant CA227838, NIH grant F30 AA024676 and philanthropic donations provided financial support for this manuscript.
We are grateful to Ms. Marsha Danley, Department of Pathology & Laboratory Medicine for technical assistance with immunohistochemistry optimization and staining.
- Patel ZM, Fernandez-Miranda J, Hwang PH, Nayak JV, Dodd R, Sajjadi H, Jackler RK. Letter: Precautions for Endoscopic Transnasal Skull Base Surgery During the COVID-19 Pandemic. Neurosurgery. 2020 Jul 1;87(1):E66-E67. doi: 10.1093/neuros/nyaa125. PMID: 32293678; PMCID: PMC7184431.
- Huang X, Zhu W, Zhao H, Jiang X. In Reply: Precautions for Endoscopic Transnasal Skull Base Surgery During the COVID-19 Pandemic. Neurosurgery. 2020 Aug 1;87(2):E160-E161. doi: 10.1093/neuros/nyaa145. PMID: 32302398; PMCID: PMC7188152.
- Zou L, Ruan F, Huang M, Liang L, Huang H, Hong Z, Yu J, Kang M, Song Y, Xia J, Guo Q, Song T, He J, Yen HL, Peiris M, Wu J. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. N Engl J Med. 2020 Mar 19;382(12):1177-1179. doi: 10.1056/NEJMc2001737. Epub 2020 Feb 19. PMID: 32074444; PMCID: PMC7121626.
- Mawaddah A, Gendeh HS, Lum SG, Marina MB. Upper respiratory tract sampling in COVID-19. Malays J Pathol. 2020 Apr;42(1):23-35. PMID: 32342928.
- Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020 Mar;579(7798):270-273. doi: 10.1038/s41586-020-2012-7. Epub 2020 Feb 3. Erratum for: Nature. 2020 Dec;588(7836):E6. PMID: 32015507; PMCID: PMC7095418.
- Sims AC, Baric RS, Yount B, Burkett SE, Collins PL, Pickles RJ. Severe acute respiratory syndrome coronavirus infection of human ciliated airway epithelia: role of ciliated cells in viral spread in the conducting airways of the lungs. J Virol. 2005 Dec;79(24):15511-24. doi: 10.1128/JVI.79.24.15511-15524.2005. PMID: 16306622; PMCID: PMC1316022.
- Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell. 2020 Apr 16;181(2):271-280.e8. doi: 10.1016/j.cell.2020.02.052. Epub 2020 Mar 5. PMID: 32142651; PMCID: PMC7102627.
- Chen M, Shen W, Rowan NR, Kulaga H, Hillel A, Ramanathan M Jr, Lane AP. Elevated ACE2 expression in the olfactory neuroepithelium: implications for anosmia and upper respiratory SARS-CoV-2 entry and replication. bioRxiv [Preprint]. 2020 May 9:2020.05.08.084996. doi: 10.1101/2020.05.08.084996. Update in: Eur Respir J. 2020 Aug 18;: PMID: 32511390; PMCID: PMC7263519.
- Bertram S, Heurich A, Lavender H, Gierer S, Danisch S, Perin P, Lucas JM, Nelson PS, Pöhlmann S, Soilleux EJ. Influenza and SARS-coronavirus activating proteases TMPRSS2 and HAT are expressed at multiple sites in human respiratory and gastrointestinal tracts. PLoS One. 2012;7(4):e35876. doi: 10.1371/journal.pone.0035876. Epub 2012 Apr 30. PMID: 22558251; PMCID: PMC3340400.
- Hamming I, Timens W, Bulthuis ML, Lely AT, Navis G, van Goor H. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J Pathol. 2004 Jun;203(2):631-7. doi: 10.1002/path.1570. PMID: 15141377; PMCID: PMC7167720.
- Jia HP, Look DC, Shi L, Hickey M, Pewe L, Netland J, Farzan M, Wohlford-Lenane C, Perlman S, McCray PB Jr. ACE2 receptor expression and severe acute respiratory syndrome coronavirus infection depend on differentiation of human airway epithelia. J Virol. 2005 Dec;79(23):14614-21. doi: 10.1128/JVI.79.23.14614-14621.2005. PMID: 16282461; PMCID: PMC1287568.
- Pan Y, Zhang D, Yang P, Poon LLM, Wang Q. Viral load of SARS-CoV-2 in clinical samples. Lancet Infect Dis. 2020 Apr;20(4):411-412. doi: 10.1016/S1473-3099(20)30113-4. Epub 2020 Feb 24. PMID: 32105638; PMCID: PMC7128099.
- Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA. 2020 Mar 17;323(11):1061-1069. doi: 10.1001/jama.2020.1585. Erratum in: JAMA. 2021 Mar 16;325(11):1113. PMID: 32031570; PMCID: PMC7042881.
- Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020 Feb 15;395(10223):497-506. doi: 10.1016/S0140-6736(20)30183-5. Epub 2020 Jan 24. Erratum in: Lancet. 2020 Jan 30;: PMID: 31986264; PMCID: PMC7159299.
- Chen R, Liang W, Jiang M, Guan W, Zhan C, Wang T, Tang C, Sang L, Liu J, Ni Z, Hu Y, Liu L, Shan H, Lei C, Peng Y, Wei L, Liu Y, Hu Y, Peng P, Wang J, Liu J, Chen Z, Li G, Zheng Z, Qiu S, Luo J, Ye C, Zhu S, Liu X, Cheng L, Ye F, Zheng J, Zhang N, Li Y, He J, Li S, Zhong N; Medical Treatment Expert Group for COVID-19. Risk Factors of Fatal Outcome in Hospitalized Subjects With Coronavirus Disease 2019 From a Nationwide Analysis in China. Chest. 2020 Jul;158(1):97-105. doi: 10.1016/j.chest.2020.04.010. Epub 2020 Apr 15. PMID: 32304772; PMCID: PMC7158802.
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.




 Save to Mendeley
Save to Mendeley
